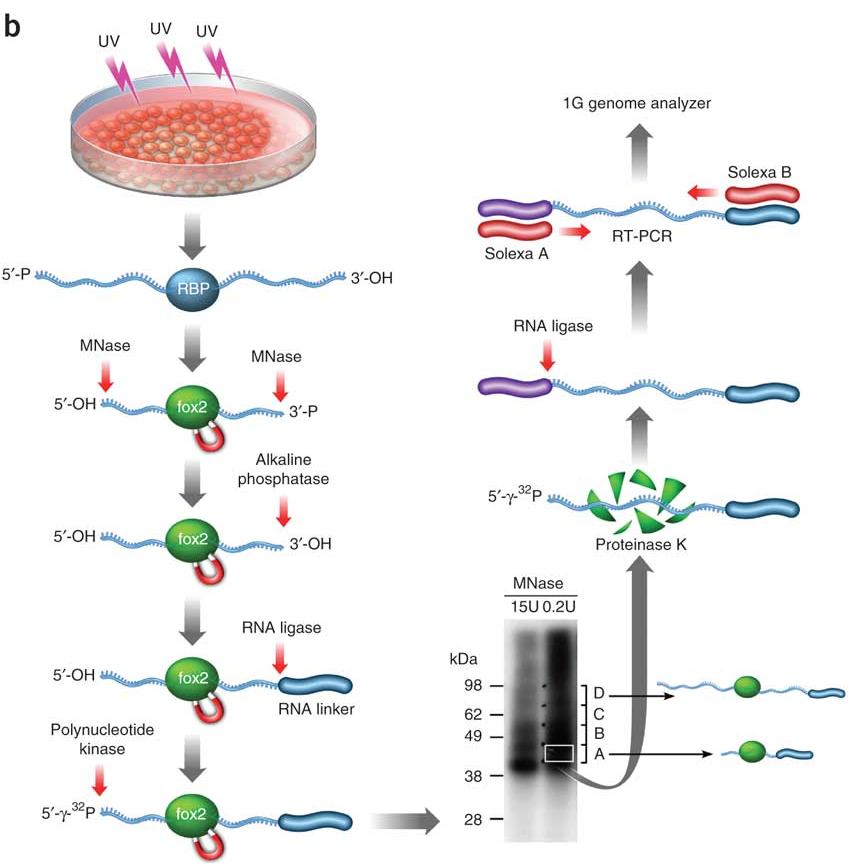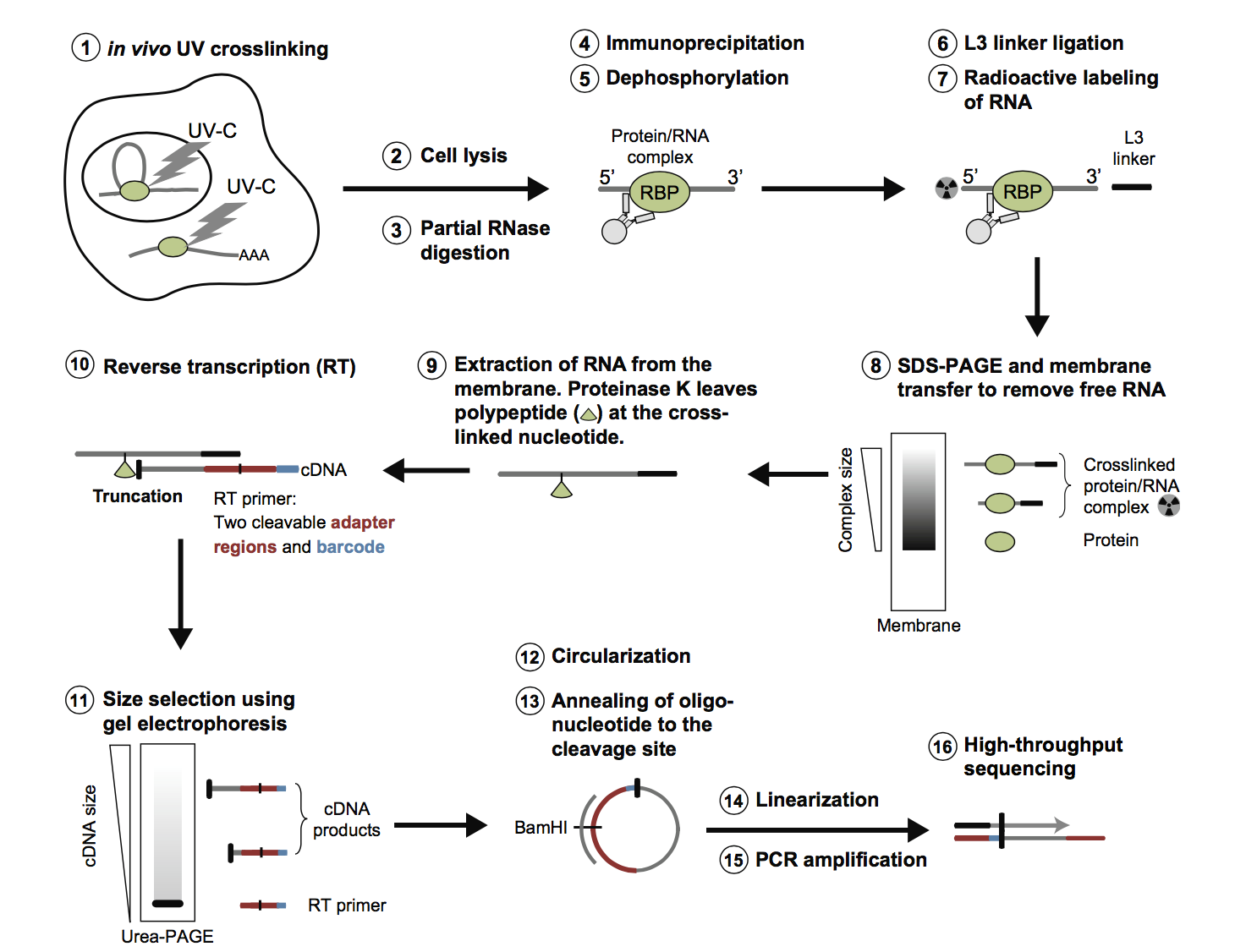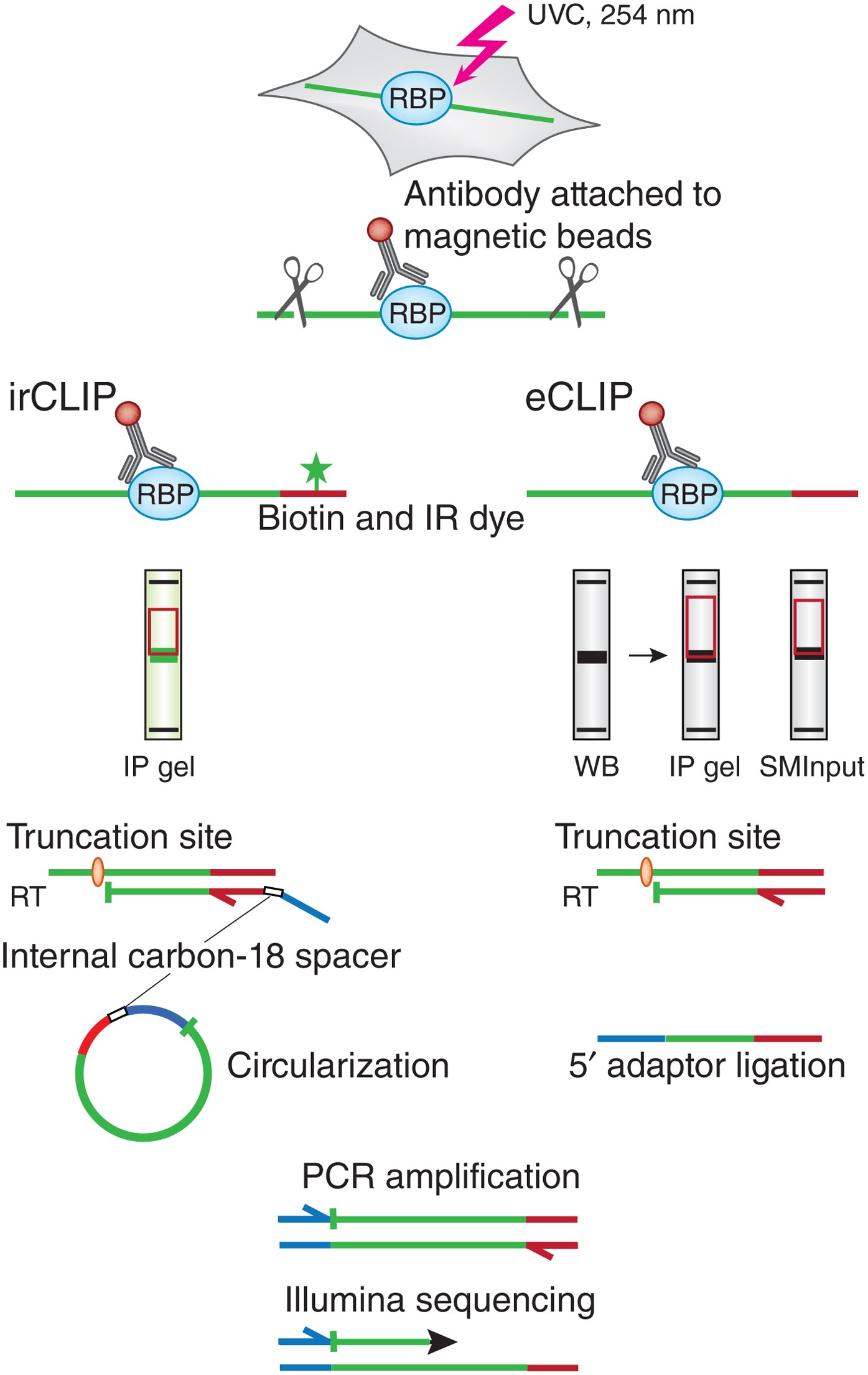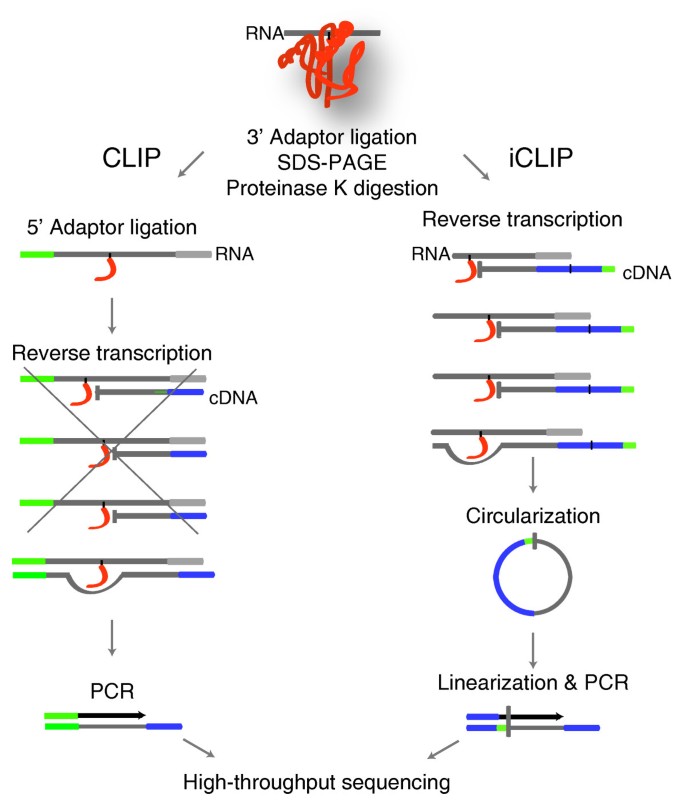
Analysis of CLIP and iCLIP methods for nucleotide-resolution studies of protein-RNA interactions | Genome Biology | Full Text

RNA-binding protein hnRNPLL regulates mRNA splicing and stability during B-cell to plasma-cell differentiation | PNAS

Optimization of PAR-CLIP for transcriptome-wide identification of binding sites of RNA-binding proteins - ScienceDirect

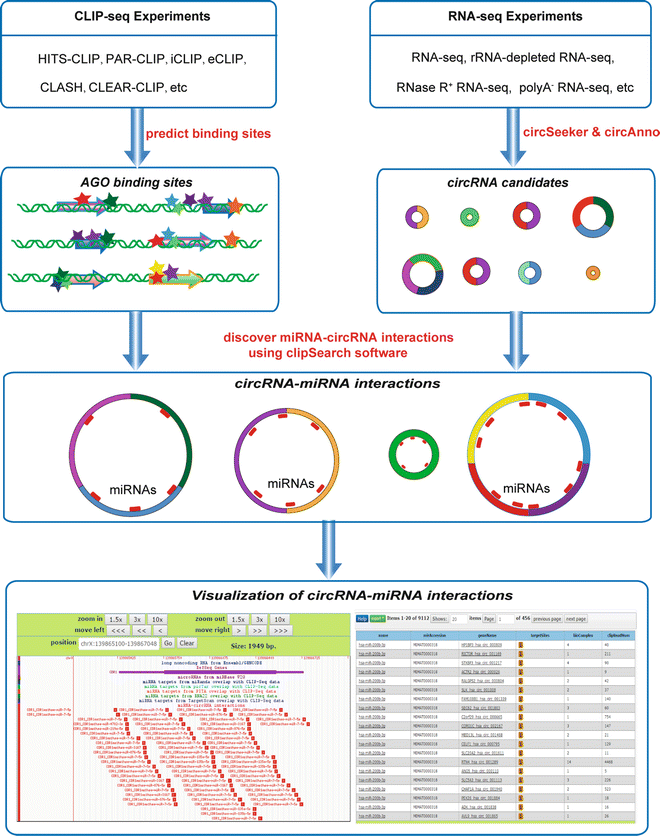
starBase v2.0: decoding Interaction Networks of lncRNAs, miRNAs, ceRNAs, RNA-Binding Proteins and mRNAs from large-scale tumor samples and CLIP-Seq (HITS-CLIP, PAR-CLIP, iCLIP, CLASH) data

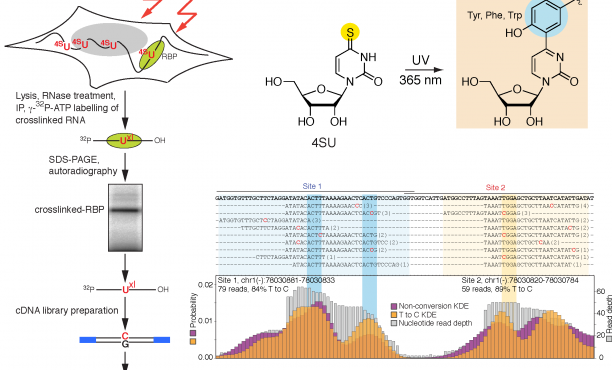
Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

Transcriptome-wide Identification of RNA-Binding Protein and MicroRNA Target Sites by PAR-CLIP: Cell

PAR-CliP - A Method to Identify Transcriptome-wide the Binding Sites of RNA Binding Proteins | Protocol

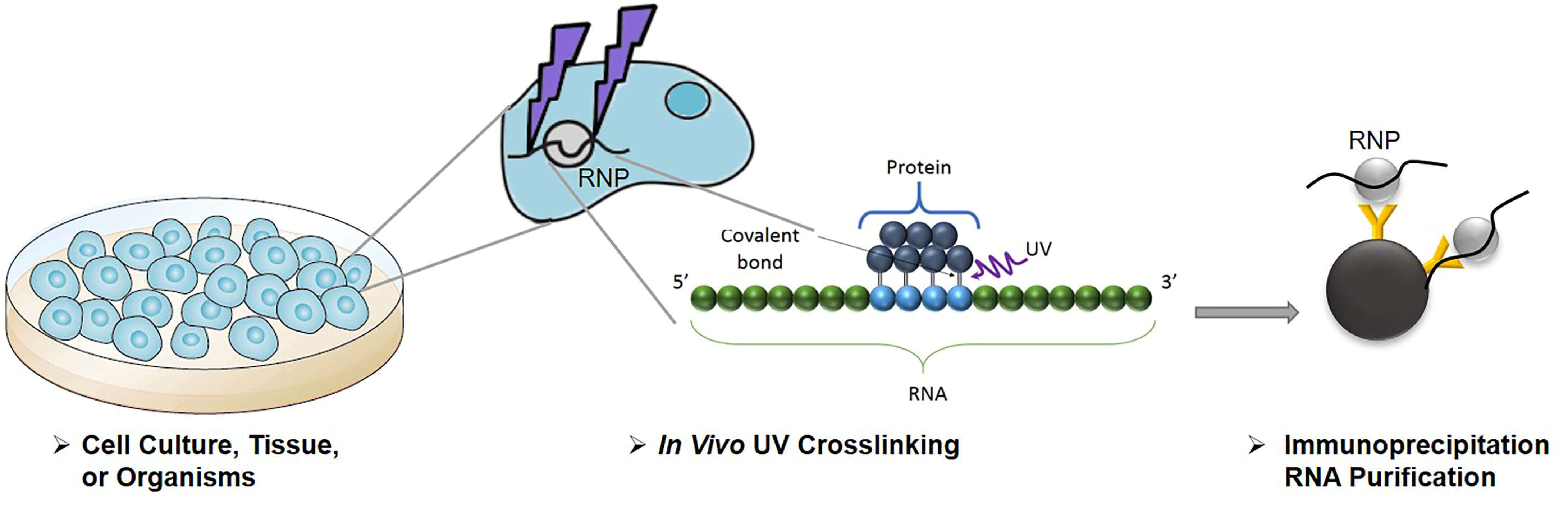
Mapping transcriptome-wide protein-RNA interactions to elucidate RNA regulatory programs. - Abstract - Europe PMC
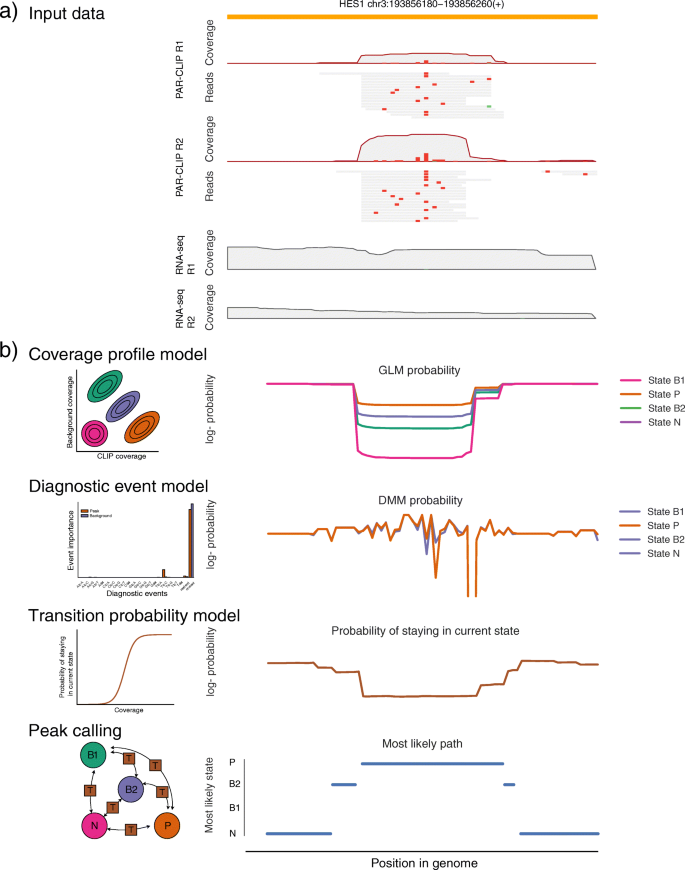
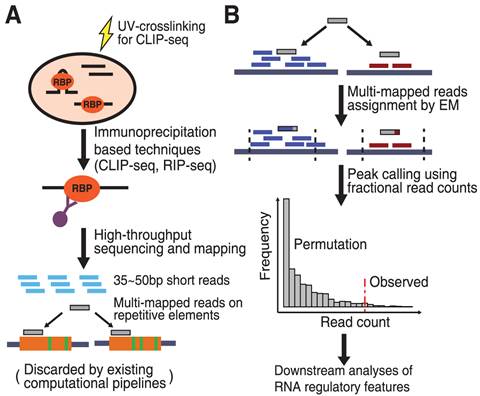
omniCLIP: probabilistic identification of protein-RNA interactions from CLIP-seq data | Genome Biology | Full Text